Microalgae are photosynthetic single-cell microorganisms capable of storing energy in the form of lipids in very large quantities. Operating this lipid production is of growing interest for numerous biotechnological applications, such as the production of biofuels. In this study, the researchers of LPCV used the oil-producing microalga
Microchloropsis gaditana as a model organism due to its ability to produce high levels of
biomass* and lipids, and its tolerance to a wide range of culture conditions (pH, temperature, salinity). In addition to having a complex metabolism, due to the unique nature of its evolutionary history,
Microchloropsis gaditana produces and accumulates lipids when the environment is deficient in nitrogen, thereby stopping its biomass production. To optimise and improve the yield of this microalga, an in silico approach has been developed: iMgadit23, a
genome-wide metabolic model (GEM)* for
Microchloropsis gaditana.
In addition to predicting
metabolic flows*, which are based on system constraints, GEMs enable the prediction of the influence of genetic and environmental factors on
cellular phenotypes*.
iMgadit23 is a new metabolic model of
Microchloropsis gaditana with a complete and validated lipid metabolism. The metabolic pathways have all been exhaustively annotated and can be visualised using a 2D ESCHER map (https://escher.github.io).
Following validation of the model—based on intrinsic quality characteristics such as reaction balance, annotations, etc., as well as experimental data—this model has enabled the simulation of the strain's behaviour in different environments and genetic conditions, and has helped elucidate the metabolic phenotype of a mutant strain exhibiting a highly interesting lipid profile (an eightfold increase in lipid content compared to the wild-type strain).
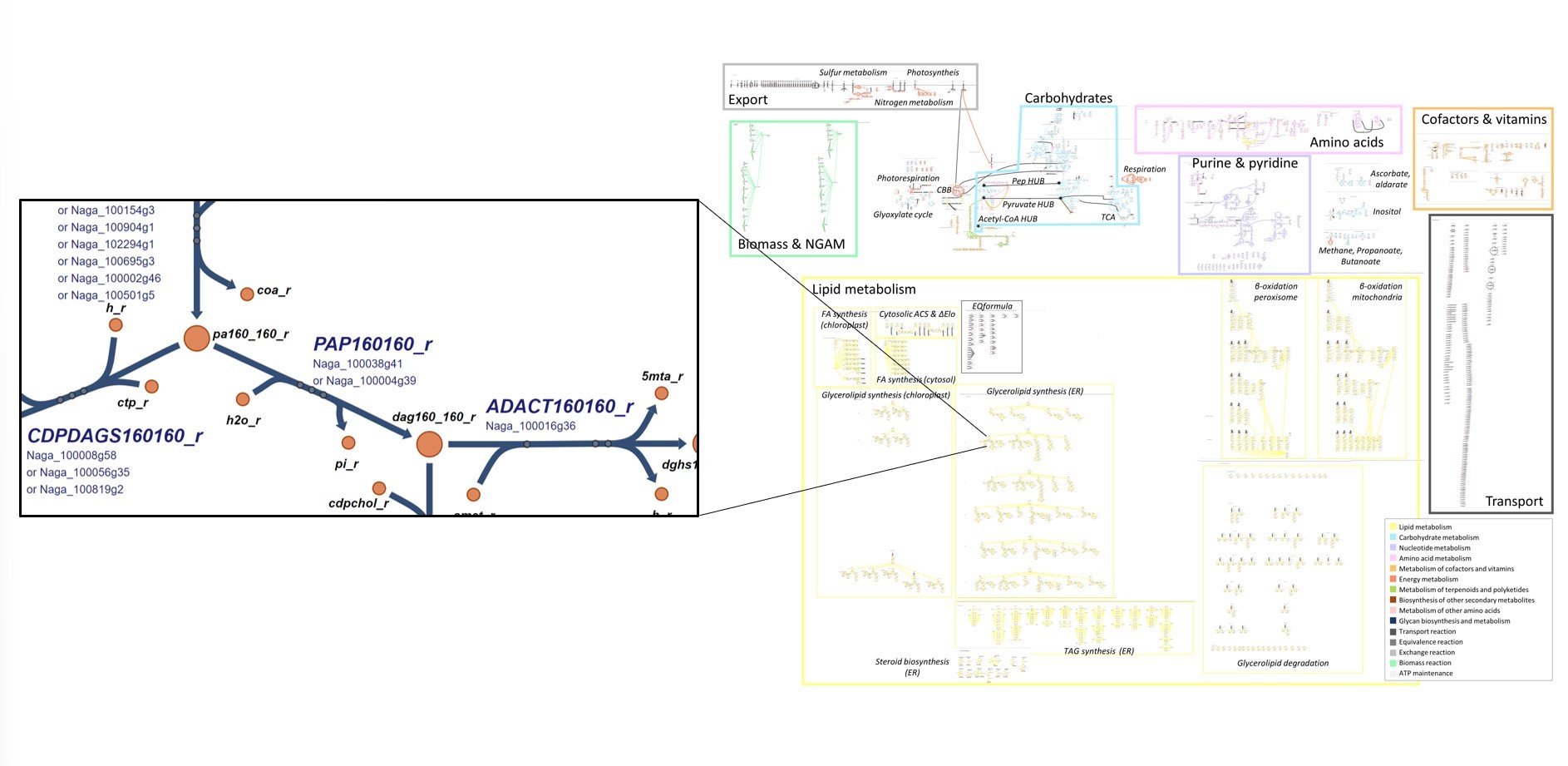
© CEA-Irig/LPCV/Lipid/J. Jouhet
Figure: Escher diagram – Visualisation of the metabolic model of Microchloropsis gaditana.
The development of this new genome-wide metabolic model, iMgadit23, makes possible, for the first time, to fully model the lipid metabolism of
Microchloropsis gaditana, thereby enabling the cost-effective elucidation of the metabolic phenotype of candidate mutant strains with a lipid profile that yields higher productivity than the wild-type strain.
As the lipid metabolism has been fully characterised, the strategy used to construct this model is intended to be applicable and transferable to any other living organism.
Lipid metabolism*: Set of chemical reactions involved in the production, conversion and storage of lipids within an organism.
Biomass*: Organic matter produced, comprising all the organic molecules (lipids, proteins, carbohydrates) that make up living matter.
Genome-wide metabolic model (GEM)*: Metabolic network bringing together all known metabolic information about a biological system: metabolites, reactions, genes, enzymes and the associated 'gene-protein-reaction' rules.
Metabolic flows*: Rate at which products are synthesised.
Cellular phenotype*: Set of observable characteristics of a cell resulting from the expression of its genes and the influence of its environment.
UMR : CEA, CNRS, INRAE, UGA
Fundings : Cifre PhD (TotalEnergies&CEA) [2020/1063] and GRAL, [ANR-17-EURE-0003]
Collaborations : LPCV, TotalEnergies (France), SilicoLife Lda (Portugal).